Department Directory
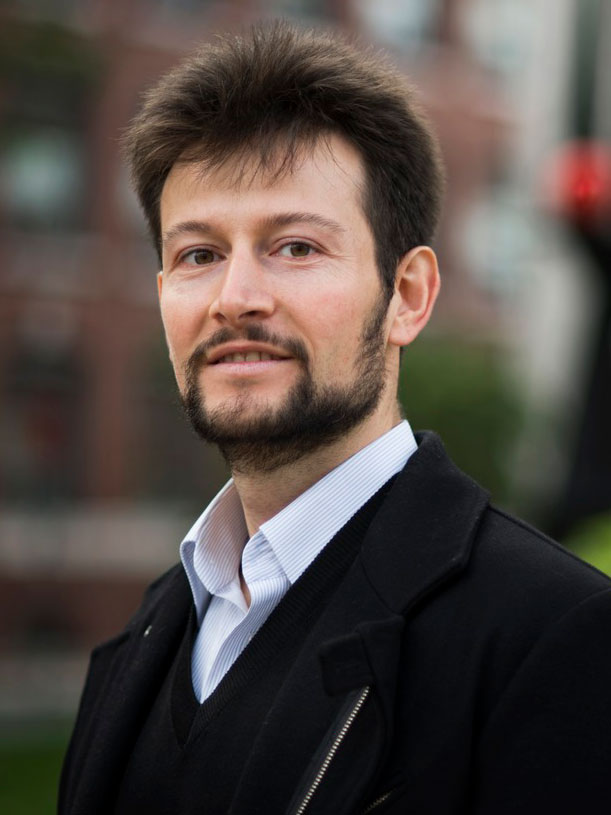

Nikolai Slavov
Associate Professor,
Bioengineering Single-cell proteomics, immunology, cancer drug resistance, translational regulation, quantitative systems biology, mass-spectrometry, macrophage polarization, neurodegenerative diseases
- n.slavov@northeastern.edu
- 617.373.6405
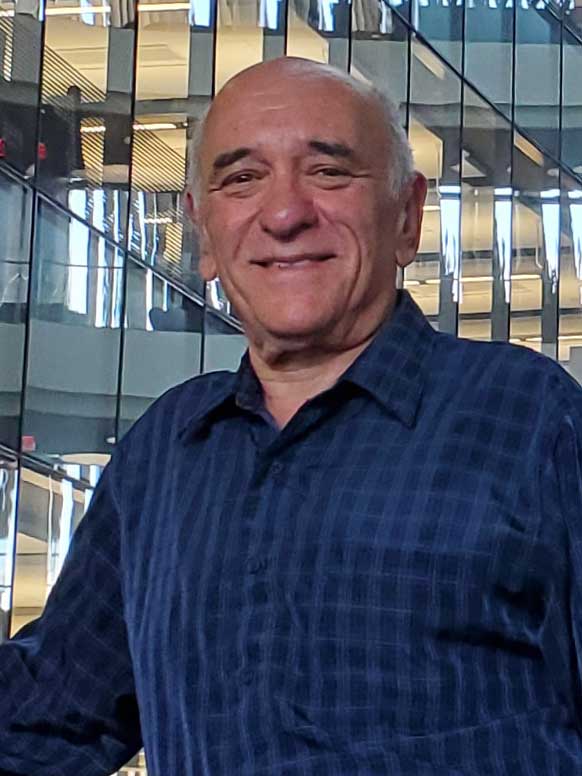

Eduardo Sontag
University Distinguished Professor,
jointly appointed in
Electrical and Computer Engineering & Bioengineering
Feedback control theory, systems biology, cancer, and biomedicine
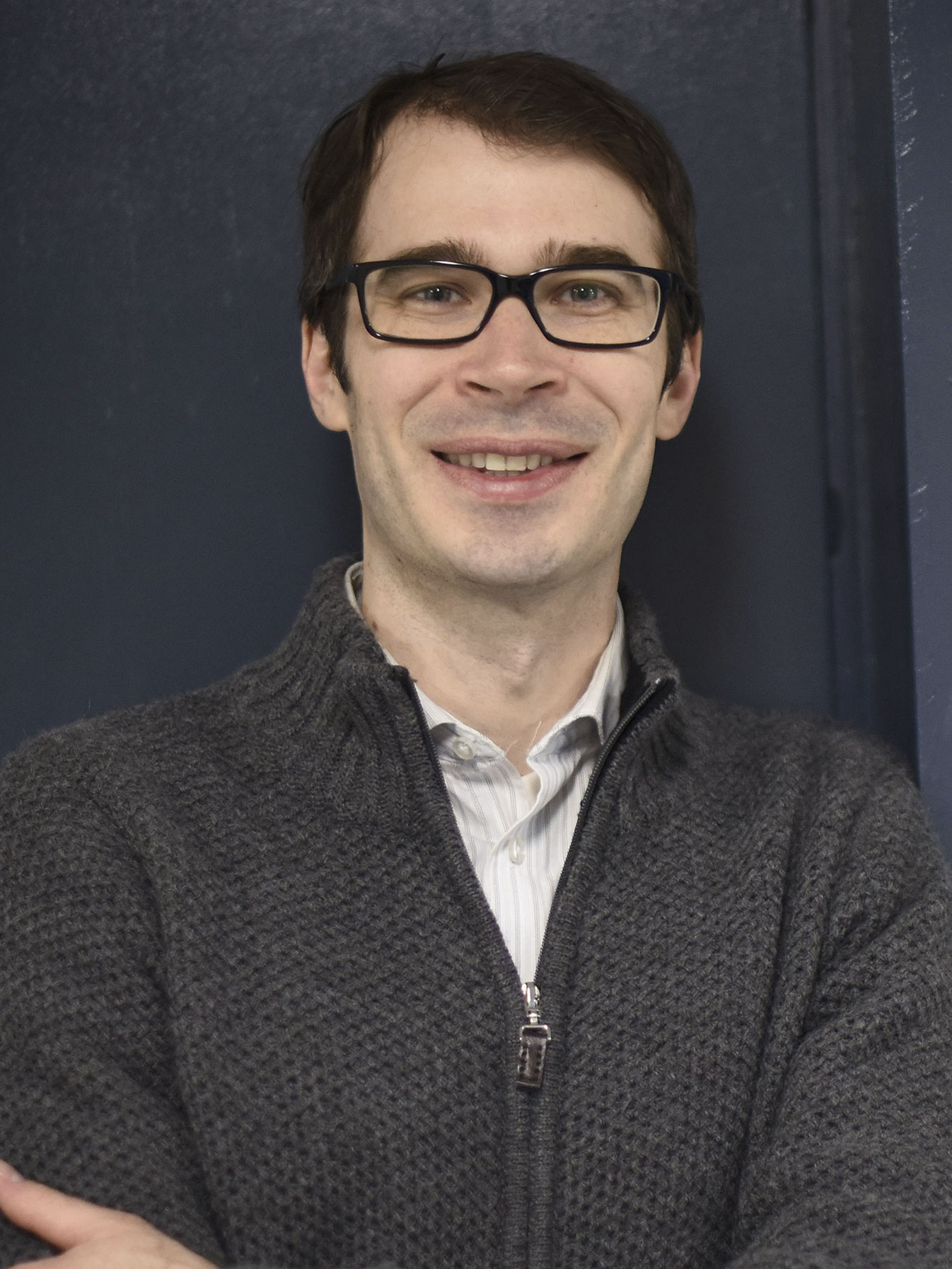

Bryan Spring
Affiliated Faculty,
Bioengineering targeted photomedicine, biophysical microscopy and cancer biology
- b.spring@northeastern.edu
- 617.373.5303
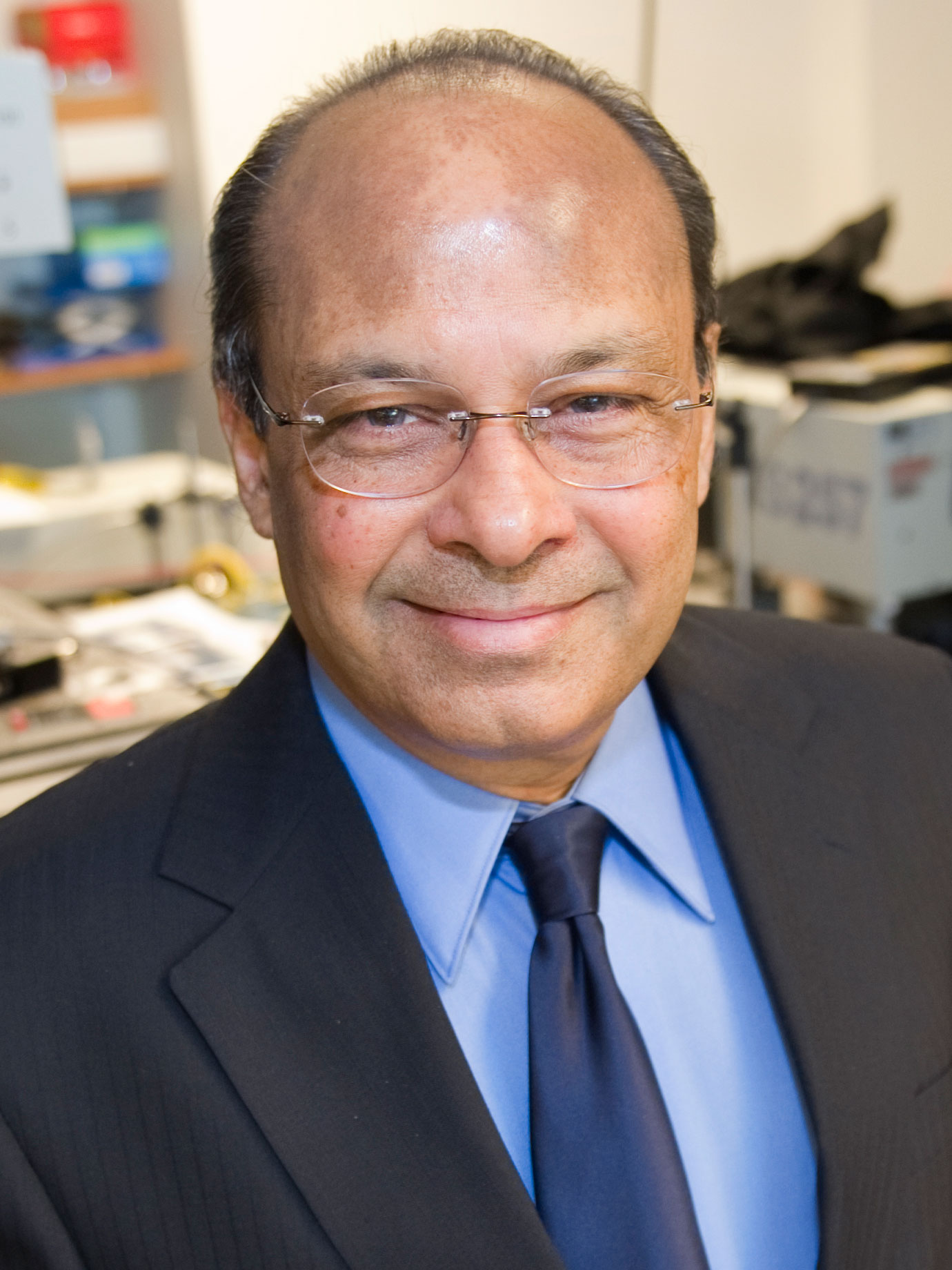

Srinivas Sridhar
Affiliated Faculty,
Bioengineering nanomedicine; neurotechnology; drug delivery, MRI imaging
- s.sridhar@northeastern.edu
- 617.373.2930

Armen Stepanyants
Affiliated Faculty,
Bioengineering Theoretical Condensed Matter And Biological Physics
- a.stepanyants@northeastern.edu
- 617.373.2944
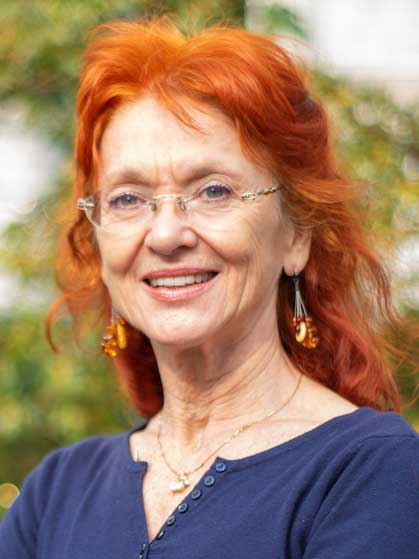

Dagmar Sternad
University Distinguished Professor,
jointly appointed in
Biology & Electrical and Computer Engineering
Motor control and learning, variability and stability, virtual rehabilitation, dynamic modeling, rhythmic and discrete movements as primitives for action
- d.sternad@northeastern.edu
- 617.373.5093

Tao Sun
Assistant Professor,
Bioengineering Focused Ultrasound, Ultrasound Imaging, Neuroimaging, Drug Delivery, Immunomodulation and Immunoengineering, Gliomblastoma, Alzheimer’s Disease

Amir Vahabikashi
Assistant Professor,
Bioengineering Cell and nucleus mechanobiology, soft bioelectronics for organoid/tissue scale mechanobiology and regenerative engineering, mechanotransduction, implantable bioelectronics

Kiran Vanaja
Assistant Research Professor,
Bioengineering Systems biology of signaling networks in cancer (and) therapeutics, metastasis and embryonic development

Jacob Walker
Associate Co-op Coordinator & Assistant Director,
Cooperative Education - jw.walker@northeastern.edu
- 617.373.5393

Lei Wang
Assistant Professor,
jointly appointed in
Bioengineering & Biology
Mammalian synthetic biology tools to advance cancer cellular therapy, regenerative medicine, and microfluidic human organ models

Ning Wang
Professor,
Bioengineering Cellular and molecular mechanobiology, mechanomedicine, and mechanohealth; cancer cell biology and mechanics; stem cell biology and mechanics; mechanomemory and mechanoresilience, mechanobiotechnologies and their applications to cells, tissues, and organisms

Meni Wanunu
Professor,
jointly appointed in
Physics & Bioengineering
Our group investigates biomolecules at the single-molecule level. We develop nanopore-based and other nanotechnology-based methods for probing the structure and dynamic behavior of biomolecules. We employ optical waveguides and single-molecule enzymatic approaches for RNA sequencing, and utilize engineered nanopore sensors for applications in single-molecule proteomics. We are experimentalists, but we also use advanced computational tools to perform big data analysis.
- m.wanunu@northeastern.edu
- 617.373.7412

Jing-Ke Weng
Professor,
jointly appointed in
Chemistry & Chemical Biology & Bioengineering
Natural product biochemistry, plant abiotic and biotic interactions, carbon sequestration, agricultural biotechnology, food allergy, drug discovery

Paul Whitford
Affiliated Faculty,
Bioengineering dynamics of large-scale molecular machines, working to identify the physical principles that guide biomolecular dynamics, using molecular simulation approaches to interpret experimental data from a wide range of techniques, including biochemical, small-angle X-ray scattering and cryogenic electron microscopy
- p.whitford@northeastern.edu
- 617.373.2952

Mark Williams
Affiliated Faculty,
Bioengineering single molecule biophysics, molecular biophysics
- ma.williams@northeastern.edu
- 617.373.5705
Rebecca Willits
Professor and Chairperson,
Chemical Engineering Mechanobiology of neural systems;
Systems for nerve regeneration;
Diversity and equity in engineering
- r.willits@northeastern.edu
- 617.373.2989



